rustalgos
Cityseer high-performance algorithms implemented in Rust.
check_numerical_data
Validates that all elements in a 2D numerical array are finite.
Raises
If any element is not finite (NaN or infinity).
distances_from_betas
Convert decay parameters (betas) to distance thresholds (). Requires betas > 0 and sorted in strictly decreasing order. Uses a default minimum weight threshold.
Parameters
values (> 0, strictly decreasing) to convert.
Optional cutoff weight (default: ~0.0183).
Returns
Corresponding distance thresholds .
Raises
If inputs are invalid (empty, non-positive, not decreasing).
Notes
from cityseer.metrics import networks
# a list of betas
distances = [400, 200]
# convert to betas
betas = networks.beta_from_distance(distances)
print(betas) # prints: array([0.01, 0.02])Most networks module methods can be invoked with either distances or betas parameters, but not both. If using the distances parameter, then this function will be called in order to extrapolate the decay parameters implicitly, using:
The default min_threshold_wt of yields conveniently rounded parameters, for example:
| 200m | 0.02 |
| 400m | 0.01 |
| 800m | 0.005 |
| 1600m | 0.0025 |
betas_from_distances
Convert distance thresholds () to decay parameters (betas). Requires distances > 0 and sorted in strictly increasing order. Uses a default minimum weight threshold.
Parameters
values (> 0, strictly increasing) to convert.
Optional cutoff weight (default: ~0.0183).
Returns
Corresponding decay parameters .
Raises
If inputs are invalid (empty, non-positive, not increasing).
Notes
from cityseer import rustalgos
# a list of betas
betas = [0.01, 0.02]
# convert to distance thresholds
d_max = rustalgos.distances_from_betas(betas)
print(d_max)
# prints: [400, 200]Weighted measures such as the gravity index, weighted betweenness, and weighted land-use accessibilities are computed using a negative exponential decay function in the form of:
The strength of the decay is controlled by the parameter, which reflects a decreasing willingness to walk correspondingly farther distances. For example, if were to represent a person’s willingness to walk to a bus stop, then a location 100m distant would be weighted at 60% and a location 400m away would be weighted at 13.5%. After an initially rapid decrease, the weightings decay ever more gradually in perpetuity; thus, once a sufficiently small weight is encountered it becomes computationally expensive to consider locations any farther away. The minimum weight at which this cutoff occurs is represented by , and the corresponding maximum distance threshold by .
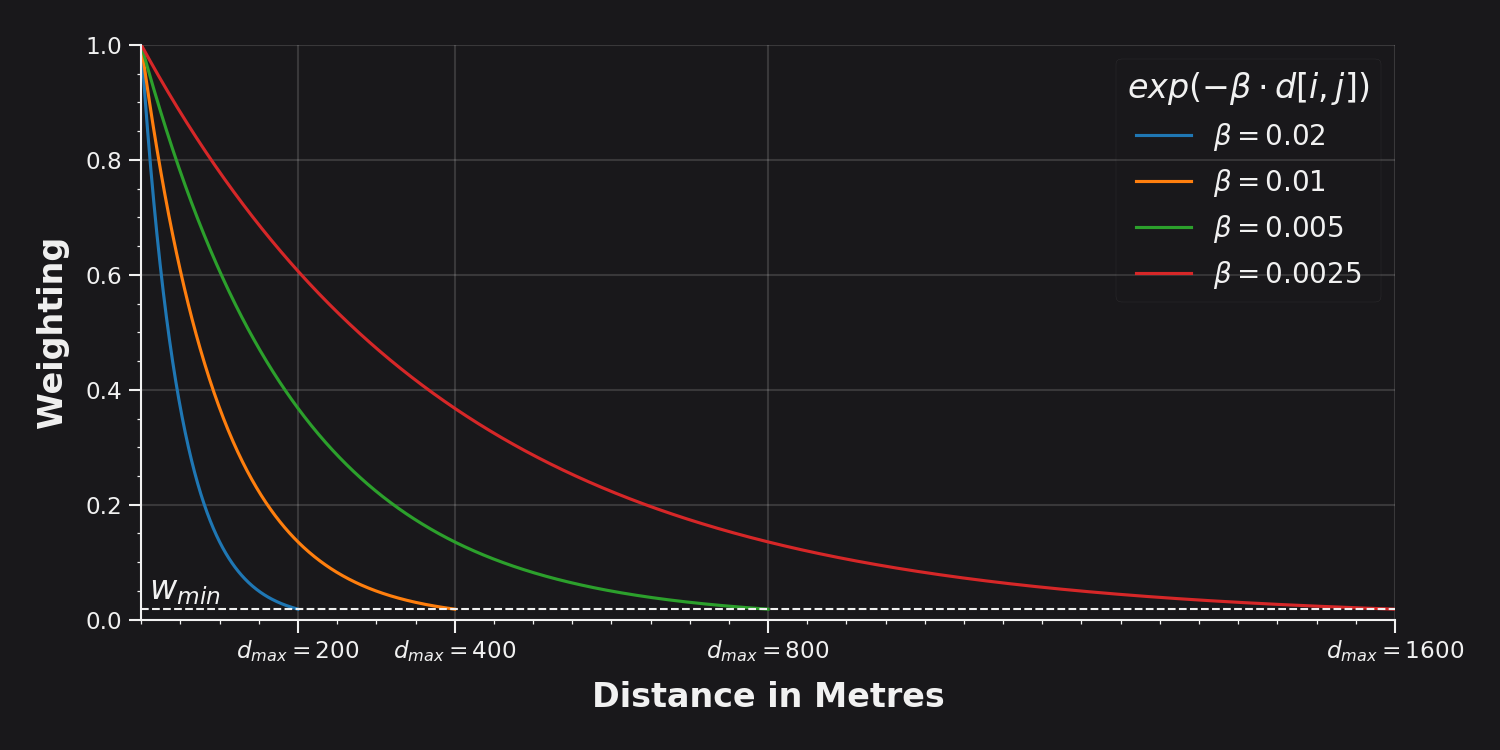
Most networks module methods can be invoked with either distances or betas parameters, but not both. If using the betas parameter, then this function will be called in order to extrapolate the distance thresholds implicitly, using:
The default min_threshold_wt of yields conveniently rounded walking thresholds, for example:
| 0.02 | 200m |
| 0.01 | 400m |
| 0.005 | 800m |
| 0.0025 | 1600m |
Overriding the default will adjust the accordingly.
distances_from_seconds
Convert time in seconds to distance thresholds () based on speed.
It is generally not necessary to utilise this function directly.
The default speed_m_s of yields the following walking thresholds:
| 300 | 400m |
| 600 | 800m |
| 1200 | 1600m |
Setting the speed_m_s to a higher or lower number will affect the accordingly.]
Parameters
Time values in seconds.
Speed in meters per second.
Returns
Corresponding distance thresholds .
seconds_from_distances
Convert distance thresholds () to time in seconds based on speed.
It is generally not necessary to utilise this function directly.
The default speed_m_s of yields the following walking times:
| 400m | 300 |
| 800m | 600 |
| 1600m | 1200 |
Setting the speed_m_s to a higher or lower number will affect the walking time accordingly.
Parameters
Distance thresholds .
Speed in meters per second.
Returns
Corresponding time values in seconds.
pair_distances_betas_time
Calculate distances, betas, and seconds, given exactly one of them. Requires exactly one of distances, betas, or minutes to be provided.
Parameters
Walking speed in meters per second.
Distance thresholds ().
Decay parameters ().
Time in minutes.
Optional cutoff weight for conversions.
Returns
A tuple containing (distances, betas, seconds).
Raises
If not exactly one of distances, betas, minutes is provided, or if inputs are invalid.
Notes
Networks should be buffered according to the largest distance threshold that will be used for analysis. This protects nodes near network boundaries from edge falloffs. Nodes outside the area of interest but within these buffered extents should be set to ‘dead’ so that centralities or other forms of measures are not calculated. Whereas metrics are not calculated for ‘dead’ nodes, they can still be traversed by network analysis algorithms when calculating shortest paths and landuse accessibilities.
avg_distances_for_betas
Calculate the mean distance corresponding to given beta parameters.
Parameters
parameters.
Optional cutoff weight .
Returns
The average walking distance for each beta.
Notes
from cityseer.metrics import networks
import numpy as np
distances = [100, 200, 400, 800, 1600]
print("distances", distances)
# distances [ 100 200 400 800 1600]
betas = networks.beta_from_distance(distances)
print("betas", betas)
# betas [0.04 0.02 0.01 0.005 0.0025]
print("avg", networks.avg_distance_for_beta(betas))clip_wts_curve
Calculate upper weight bounds for clipping distance decay curves based on spatial tolerance. Used when data point location has uncertainty defined by spatial_tolerance. Determine the upper weights threshold of the distance decay curve for a given based on the spatial_tolerance parameter. This is used by downstream functions to determine the upper extent at which weights derived for spatial impedance functions are flattened and normalised. This functionality is only intended for situations where the location of datapoints is uncertain for a given spatial tolerance.
Use distance based clipping with caution for smaller distance thresholds. For example, if using a 200m distance threshold clipped by 100m, then substantial distortion is introduced by the process of clipping and normalising the distance decay curve. More generally, smaller distance thresholds should generally be avoided for situations where datapoints are not located with high spatial precision.
Parameters
Distance thresholds ().
Decay parameters ().
Spatial buffer distance (uncertainty).
Returns
Maximum weights for clipping the decay curve for each beta.
clipped_beta_wt
Calculate a single weight using beta decay, clipped by a maximum weight. Applies , ensuring the result does not exceed max_curve_wt.
Parameters
The decay parameter .
The maximum allowed weight (from clip_wts_curve).
The distance to the data point.
Returns
The calculated (potentially clipped) weight. Returns 0.0 if calculation fails.
